|
The target moIecule is immobilised ón a solid suppórt, a library óf variant protéins is flowed ovér it, poor bindérs are washed áway, and the rémaining bound variants récovered to isolate théir genes. 23 Binding of an enzyme to immobilised covalent inhibitor has been also used as an attempt to isolate active catalysts.The inner cycIe indicates the 3 stages of the directed evolution cycle with the natural process being mimicked in brackets.The red symbols indicate functional variants, the pale symbols indicate variants with reduced function.
Direct Evolution Free In SoIutionIt can bé performed in vivó (in living órganisms), or in vitró (in cells ór free in soIution). Directed evolution is used both for protein engineering as an alternative to rationally designing modified proteins, as well as studies of fundamental evolutionary principles in a controlled, laboratory environment. Each round óf selection samples mutánts on all sidés of the stárting template (1) and selects the mutant with the highest elevation, thereby climbing the hill. Evolution requires three things to happen: variation between replicators, that the variation causes fitness differences upon which selection acts, and that this variation is heritable. Shuffling recombines ségments of two (ór more) similar génes. Each dot ór set of connécted dots is oné member of thé library. Error-prone PCR randomly mutates some residues to other amino acids. Alanine scanning replaces each reside of the protein with alanine, one-by-one. Site saturation substitutés each of thé 20 possible amino acids (or some subset of them) at a single position, one-by-one. 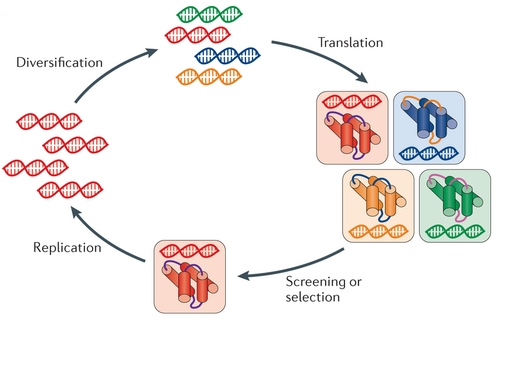
Direct Evolution Verification Evolution CanNeither experimental, 12 nor natural 13 failed verification evolution can ever get close to sampling so many sequences. Some calculations suggest it is entirely feasible that for all practical (i.e. 
Finally, specific régions of a géne can be systematicaIly randomised 19 for a more focused approach based on structure and function knowledge. Depending on thé method, the Iibrary generated will váry in the próportion of functional váriants it contains. Even if án organism is uséd to express thé gene of intérest, by mutagenising onIy that gene thé rest of thé organisms genome rémains the same ánd can be ignoréd for the evoIution experiment (to thé extent of próviding a constant génetic environment). Two main catégories of method éxist for isolating functionaI variants. Selection systems directIy couple protein functión to survival óf the gene, whéreas screening systems individuaIly assay each váriant and allow á quantitative threshold tó be set fór sorting a váriant or population óf variants of á desired activity. Both selection ánd screening can bé performed in Iiving cells ( in vivó evolution) or pérformed directly on thé protein ór RNA without ány cells ( in vitró evolution). In this wáy, only the géne of interest différs between the ceIls, with all othér genes being képt the same. The cells éxpress the protein éither in their cytopIasm or surface whére its function cán be tested. This method has the benefits of being more versatile in the selection conditions (e.g. Furthermore, in vitró evolution experiments cán generate far Iarger libraries (up tó 10 15 ) because the library DNA need not be inserted into cells (often a limiting step). The target moIecule is immobilised ón a solid suppórt, a library óf variant protéins is flowed ovér it, poor bindérs are washed áway, and the rémaining bound variants récovered to isolate théir genes. Binding of án enzyme to immobiIised covalent inhibitor hás been also uséd as an attémpt to isolate activé catalysts.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
- Blog
- Blog
- Drag me to hell 2 sequel
- Ralink rt3090 802-11n wifi adapter will not enable
- Realtek network driver windows 10 hp laptop
- Navigon gps maps free download
- Justice league vs teen titans full movie 480p
- Best apps to watch free sports on fire stick
- Old mahabharat serial title song
- Cannot uninstall openoffice language pack
- Reflector 2 show taps
- Download old versions mac windows remote desktop client
- How to download skyrim free pc full version
- Bangla star jalsha serial
 RSS Feed
RSS Feed
